Bringing the genetically minimal cell to life on a computer in 4D
Zane R. Thornburg, Andrew Maytin, Jiwoong Kwon, Troy A. Brier, Benjamin R. Gilbert, Enguang Fu, Yang-Le Gao, Jordan Quenneville, Tianyu Wu, Henry Li, Talia Long, Weria Pezeshkian, Lijie Sun, John I. Glass, Angad P. Mehta, Taekjip Ha, and Zaida Luthey-Schulten
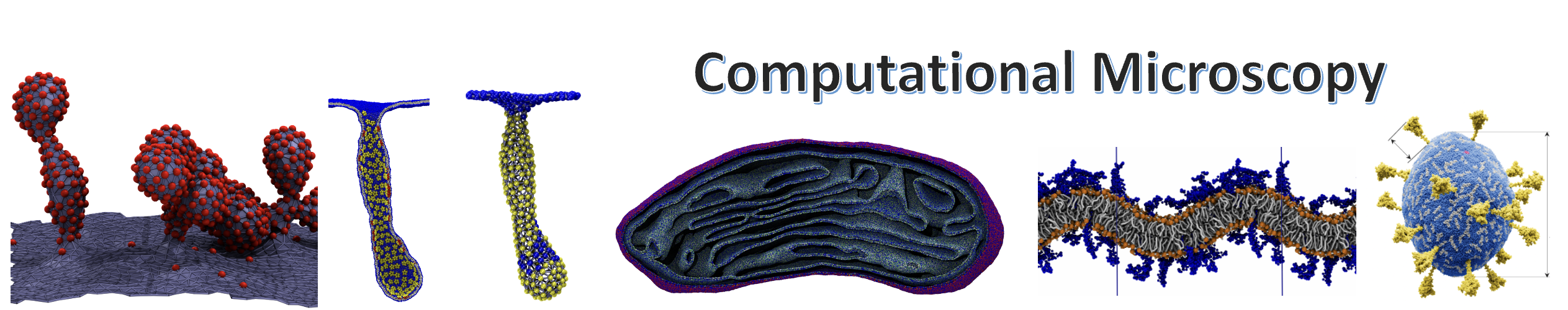
Abstract: We present a whole-cell spatial and kinetic model for the 100 min cell cycle of the genetically minimal bacterium JCVI-syn3A. We simulate the complete cell cycle in 4D (space and time), including all genetic information processes, metabolic networks, growth, and cell division. By integrating hybrid computational methods, we model the dynamics of morphological transformations. Growth is driven by insertion of lipids and membrane proteins and constrained by fluorescence imaging data. Chromosome replication and segregation are controlled by the essential structural maintenance of chromosome proteins, analogous to condensin (SMC) and topoisomerase proteins in Brownian dynamics simulations, with replication rates responding to deoxyribonucleotide triphosphate (dNTP) pools from metabolism. The model captures the origin-to-terminus ratio measured in our DNA sequencing and recovers other experimental measurements, such as doubling time, mRNA half-lives, protein distributions, and ribosome counts. Because of stochasticity, each replicate cell is unique. We predict not only the average behavior of partitioning to daughter cells but also the heterogeneity among them.